|
6/30/2023 0 Comments Designing primers using amplifx 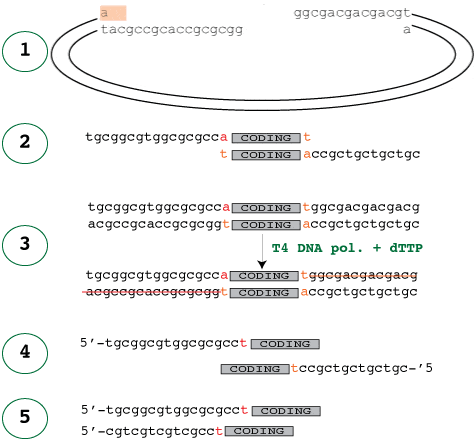 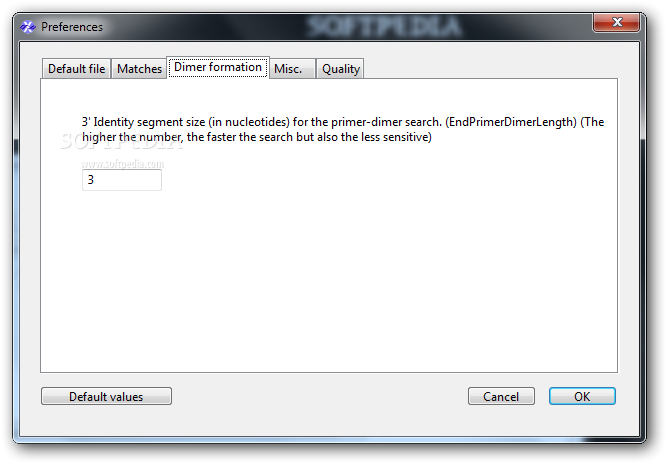
Keyboard controls: Space bar - toggle play/pause Right and Left Arrow - seek the video forwards and back Up and Down Arrow - increase and decrease the volume M key - toggle mute/unmute F key - toggle fullscreen off and on. Display Primer-BLAST results in the context of the template or genome using graphical displays.Choose the best background BLAST database to check primer specificity or identify targets.Interpret primer alignments on intended and off-target matches to pick the best primers.Identify potential products for a pair of primers.Design primers for specific genomic templates such as an exon of a gene.Design primers specific for mRNA (cDNA) templates that will not cross amplify genomic DNA.Primer-BLAST primers are suitable for use in all PCR based molecular biology protocols including target identification / verification, cloning, variant analysis, and gene expression. NCBI’s Primer-BLAST combines the primer design features of the popular Primer3 package with a specificity check that uses nucleotide BLAST to search for target and non-target matches in a background database. In addition, PCR amplification with specially designed primers is sometimes used to identify an organism or group of organisms based on targeted RNA or genomic DNA amplification of an isolate. Training materials from this event are available on this page.īiological researchers often design specific PCR primers to amplify a single genomic or mRNA template or a set of closely related templates. Avoid runs of 3 or more Cs or Gs at the 3 end of. Avoid mismatches between the 3 end of the primer and the template. On March 15, 2022, the NCBI Education Team provided the workshop: Using NCBI’s Primer-BLAST to Design and Analyze PCR Primers. The following points should be considered when designing PCR primers and are common to all types of PCR: T m calculation: 2☌ x (A+T) + 4☌ x (G+C) Avoid complementarity in the 23 bases at the 3 end of the primer pairs. A more recent version of this workshop is available at 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
